Q: What does "Undetermined" mean in my results?
A: "Undetermined" means that the fluorescent signal never passed the threshhold level, even after the maximum number of PCR cycles (40). In other words, that particular miRNA is not present in an amount that is detectable.
Q: Why do so many assays come up "Undetermined"? Is something wrong?
A: The panel includes many miRNA assays, of which about 20% - 40% are expressed in any given sample. A given miRNA may be expressed in some tissues, but not in others, or at certain developmental stages, or under specific conditions.
If virtually all of your assays come up undetermined, even the endogenous controls, then there is a problem. A poor quality sample is the most likely cause. Perhaps there is not enough template, or there is some contaminant that inhibits PCR. We try to screen for low-template samples, but our methods are not perfect. To guard against the possibility that something isn't working correctly in our lab, we run a positive control plate with each batch (of RT and preamp reactions), which should let us know if there is a problem with our processing or reagents.
Q: Do I need to use the SDS software? It seems like there is a lot to learn.
A: Most researchers should be able to work without the SDS software. We've prepared a spreadsheet with CT values for each assay. You can compute ΔCT values, ΔΔCT values and RQ (relative quantity) values using simple math:
ΔCT (test) = CT (test assay) - CT (test endogenous control)
ΔCT (calibrator) = CT (calibrator assay) - CT (calibrator endogenous control)
ΔΔCT = ΔCT (test) - ΔCT (calibrator)
-ΔΔCT
RQ = 2.0
Advanced users may want to use the SDS software to see amplification plots, or to adjust the analysis parameters, but we don't expect that researchers who are not already familiar with the software will get involved with it, initially.
Q: This seems a lot like the qRT/PCR that we do in our lab. Is it the same?
A: The "megaplex" reverse transcription reaction is unique to this assay, and the "pre-amplification" reaction is not frequently employed in assays other than this one. However, the basic chemistry, physics, and data analysis are the same.
Q: I need my results right away. Can you do my project first?
A: In the interest of fairness and efficiency, all projects will be processed in the order that they are received (Completed application form and all samples). We will do our best to process samples as quickly as possible.
Q: Why do I get different CT values for hsa-miR-126 and hsa-miR-126* ? Shouldn't they be the same?
A: While * and non-* miRNA's are usually related by being formed from opposite sides of the same immature miRNA hairpin there is no biological reason to assume that they will be expressed identically. In addition, these miRNA's have different sequences, and their expression is quantified using different assays which might have different sensitivities. The number following the assay name in your data is the ABI part number for the assay, and if you look at your data carefully you will see that these part numbers are not identical.
Q: What if I am informed that my RNA is not of ideal quality?
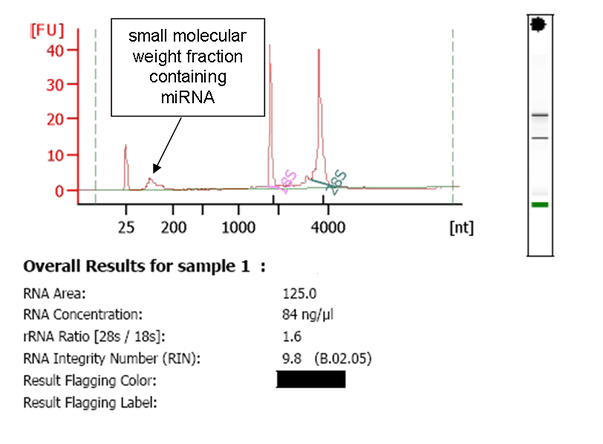
A: An aliquot of each RNA sample that has been submitted for miRNA analysis is evaluated by capillary electrophoresis on an Agilent Bioanalyzer 2100. We have found that this instrument provides a quality and concentration assessment that is a better predictor of the quality of TLDA miRNA results than is a traditional spectrophotometer. Below is an example of Bioanalyzer data from an undegraded RNA sample that contains a sufficient quantity of RNA:
A: "Undetermined" means that the fluorescent signal never passed the threshhold level, even after the maximum number of PCR cycles (40). In other words, that particular miRNA is not present in an amount that is detectable.
Q: Why do so many assays come up "Undetermined"? Is something wrong?
A: The panel includes many miRNA assays, of which about 20% - 40% are expressed in any given sample. A given miRNA may be expressed in some tissues, but not in others, or at certain developmental stages, or under specific conditions.
If virtually all of your assays come up undetermined, even the endogenous controls, then there is a problem. A poor quality sample is the most likely cause. Perhaps there is not enough template, or there is some contaminant that inhibits PCR. We try to screen for low-template samples, but our methods are not perfect. To guard against the possibility that something isn't working correctly in our lab, we run a positive control plate with each batch (of RT and preamp reactions), which should let us know if there is a problem with our processing or reagents.
Q: Do I need to use the SDS software? It seems like there is a lot to learn.
A: Most researchers should be able to work without the SDS software. We've prepared a spreadsheet with CT values for each assay. You can compute ΔCT values, ΔΔCT values and RQ (relative quantity) values using simple math:
ΔCT (test) = CT (test assay) - CT (test endogenous control)
ΔCT (calibrator) = CT (calibrator assay) - CT (calibrator endogenous control)
ΔΔCT = ΔCT (test) - ΔCT (calibrator)
-ΔΔCT
RQ = 2.0
Advanced users may want to use the SDS software to see amplification plots, or to adjust the analysis parameters, but we don't expect that researchers who are not already familiar with the software will get involved with it, initially.
Q: This seems a lot like the qRT/PCR that we do in our lab. Is it the same?
A: The "megaplex" reverse transcription reaction is unique to this assay, and the "pre-amplification" reaction is not frequently employed in assays other than this one. However, the basic chemistry, physics, and data analysis are the same.
Q: I need my results right away. Can you do my project first?
A: In the interest of fairness and efficiency, all projects will be processed in the order that they are received (Completed application form and all samples). We will do our best to process samples as quickly as possible.
Q: Why do I get different CT values for hsa-miR-126 and hsa-miR-126* ? Shouldn't they be the same?
A: While * and non-* miRNA's are usually related by being formed from opposite sides of the same immature miRNA hairpin there is no biological reason to assume that they will be expressed identically. In addition, these miRNA's have different sequences, and their expression is quantified using different assays which might have different sensitivities. The number following the assay name in your data is the ABI part number for the assay, and if you look at your data carefully you will see that these part numbers are not identical.
Q: What if I am informed that my RNA is not of ideal quality?
A: An aliquot of each RNA sample that has been submitted for miRNA analysis is evaluated by capillary electrophoresis on an Agilent Bioanalyzer 2100. We have found that this instrument provides a quality and concentration assessment that is a better predictor of the quality of TLDA miRNA results than is a traditional spectrophotometer. Below is an example of Bioanalyzer data from an undegraded RNA sample that contains a sufficient quantity of RNA:
1. RNA concentration
Our assay prescribes an initial input of 60ng total RNA per reverse transcription (RT) reaction, and a total of 120ng for the two reactions needed per sample. To achieve this initial input, we need a minimum concentration of 20ng/ul. We are not able to make up for more dilute samples by decreasing water in the reaction, as there is very little water in the reaction components. We have concluded from our pilot studies that 60ng initial RNA input per RT reaction is optimal because it amplifies the highest number of miRNA targets generating Ct values in a reliable range (Ct < 29.0) without saturating any reactions. Typically, for a human A card, the number of miRNA targets amplifying in this range is around 120. We have found that less than 60ng of input RNA will likely yield fewer targets with Ct values in this range. However, we have found that for those targets that do amplify, the data are just as reliable. In summary, if your RNA is at a low concentration, you will likely have fewer, but not poorer, data.
2. RNA quality
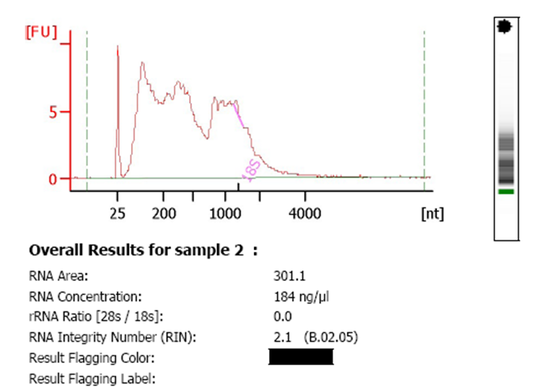
A common problem with submitted RNA is degradation. An example of degraded RNA is below:
Our assay prescribes an initial input of 60ng total RNA per reverse transcription (RT) reaction, and a total of 120ng for the two reactions needed per sample. To achieve this initial input, we need a minimum concentration of 20ng/ul. We are not able to make up for more dilute samples by decreasing water in the reaction, as there is very little water in the reaction components. We have concluded from our pilot studies that 60ng initial RNA input per RT reaction is optimal because it amplifies the highest number of miRNA targets generating Ct values in a reliable range (Ct < 29.0) without saturating any reactions. Typically, for a human A card, the number of miRNA targets amplifying in this range is around 120. We have found that less than 60ng of input RNA will likely yield fewer targets with Ct values in this range. However, we have found that for those targets that do amplify, the data are just as reliable. In summary, if your RNA is at a low concentration, you will likely have fewer, but not poorer, data.
2. RNA quality
A common problem with submitted RNA is degradation. An example of degraded RNA is below:
However, our experience is that the miRNA fraction of the total RNA is quite resistant to degradation because of its small size. Often, even degraded total RNA yields good results in the miRNA profiling assay. Similarly to issues with RNA concentration, it will be the number of targets amplifying in the reliable range that could potentially be reduced by RNA degradation rather than the quality of the data themselves. As each sample is different, we cannot provide RNA quality/quantity guidelines that will guarantee a good outcome. If any of your samples look suspect on the bioanalyzer, MDL staff will contact you before proceeding with your analysis. If you choose at that point to drop out any samples, you will be charged $7 per sample ($12 per sample outside DFCI) for the quality control assessment.
HOME | SERVICES | APPLICATION FOR SERVICES | CONTACT
©2021 Molecular Diagnostics Laboratory at Dana-Farber Cancer Institute
©2021 Molecular Diagnostics Laboratory at Dana-Farber Cancer Institute